Next: Inverse Problem Up: Independent Component Analysis For Previous: Introduction
Forward Problem
The EEG forward problem can be stated as follows: given the positions, orientations and magnitudes of dipole current sources, as well as the geometry and electrical conductivity of the head volume,and Neumann boundary conditions on the scalp:
![]()
where ![]() is a conductivity tensor and
is a conductivity tensor and ![]() are the volume currents density due to current dipoles placed within the
head. The unknown
are the volume currents density due to current dipoles placed within the
head. The unknown ![]() is the electric potential created in the head by the distribution of current
from the dipole sources. An ideal current dipole source can be described
as two point sources of opposite polarity with infinitely large current
density
is the electric potential created in the head by the distribution of current
from the dipole sources. An ideal current dipole source can be described
as two point sources of opposite polarity with infinitely large current
density ![]() and infinitely small separation d:
and infinitely small separation d:
![]()
and ![]() , the dipole strength.
, the dipole strength.
To solve Poisson's equation numerically, we began with the construction of a computational model. The realistic head geometry was obtained from MRI data, where the volume was segmented and each tissue material was labeled in the underlying voxels [23]. The segmented head volume was then tetrahedralized via a mesh generator that preserved the classification when mapping from voxels to elements [24]. For each tissue classification, we assigned a conductivity tensor from the literature [25]. A cut-through of the classified mesh is shown in Fig. 4.
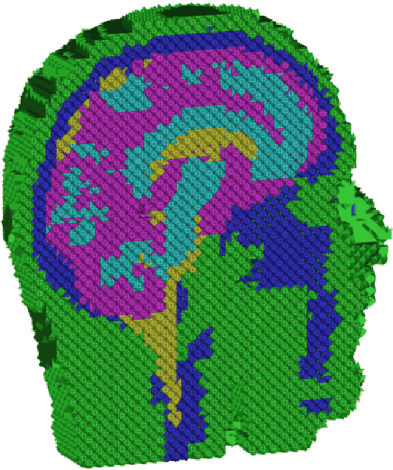
Figure 4: Cut-through of the tetrahedral mesh, with elements
colored according to conductivity classification. Green elements correspond
to skin, blue to skull, yellow to cerebro-spinal fluid, purple to gray
matter, and light blue to white matter.
We then used the finite element method (FEM) to compute a solution within the entire volume domain [26]. The FEM allows us to capture the anisotropy of conductivity and accurate boundaries of the volume. The main idea behind the FEM is to reduce a continuous problem with infinitely many unknown field values to a finite number of unknowns by discretizing the solution region into elements. Then the values of the field at any point can be approximated by interpolation functions within every element in terms of the field values at specified points called nodes. Nodes are located at the element vertices where adjacent elements are connected. Details of the FEM method can be found in [26, 27, 28].
In our study, we use tetrahedral elements and linear interpolation functions
within each tetrahedron. Our head model consists of approximately 768,000
elements and N=164,000 nodes. Once we have a geometric model, we
can assemble the matrix equations (build the matrix ![]() ) for relating field values at different nodes. This can be done using,
for example, a Rayleigh-Ritz or Galerkin method [28].
Finally, we impose boundary conditions and apply source currents. These
boundary and source conditions are incorporated within the right hand side
(RHS) of the system (vector
) for relating field values at different nodes. This can be done using,
for example, a Rayleigh-Ritz or Galerkin method [28].
Finally, we impose boundary conditions and apply source currents. These
boundary and source conditions are incorporated within the right hand side
(RHS) of the system (vector ![]() ). As a result, when we move sources we do not have to rebuild the mesh
or the matrix
). As a result, when we move sources we do not have to rebuild the mesh
or the matrix ![]() . We note that for linear interpolation functions, the RHS vector is not
sensitive to the position of a source within an element; that is, for any
position (though not orientation) within a particular tetrahedron, the
contribution to the right hand side vector is the same. This ambiguity
is relevant because it will restrict the accuracy of our inverse solution
when we attempt to recover the exact source positions.
. We note that for linear interpolation functions, the RHS vector is not
sensitive to the position of a source within an element; that is, for any
position (though not orientation) within a particular tetrahedron, the
contribution to the right hand side vector is the same. This ambiguity
is relevant because it will restrict the accuracy of our inverse solution
when we attempt to recover the exact source positions.
Using the FEM, we obtain the linear system of equations:
![]()
where ![]() is an
is an ![]() stiffness matrix,
stiffness matrix, ![]() is a source vector and
is a source vector and ![]() is the vector of unknown potentials at the nodes within the head volume.
The
is the vector of unknown potentials at the nodes within the head volume.
The ![]() matrix is sparse (containing approximately two million non-zeroes entries),
symmetric, and positive definite.
matrix is sparse (containing approximately two million non-zeroes entries),
symmetric, and positive definite.
The solution of this linear system was computed using a parallel conjugate gradient (CG) method and required approximately 12 seconds of wall-clock time on a 14 processor SGI Power Onyx with 195 MHz MIPS R10000 processors. The solution to a radially-oriented single dipole source forward problem is visualized in Fig. 5. In this image, we display an equipotential surface in wire frame, indicating the dipole location with red and blue spheres, cut-through the initial MRI data with orthogonal planes, and render the surface potential map of the bioelectric field on the cropped scalp surface.
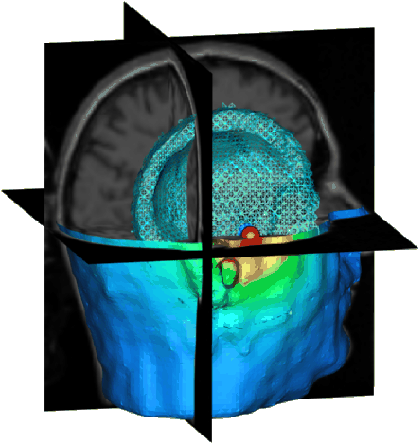
Figure 5: Solution to a single dipole source forward problem.
The underlying model is shown in the MRI planes, the dipole source is indicated
with the red and blue spheres, and the electric field is visualized by
a cropped scalp potential mapping and a wire-frame equipotential isosurface.
In order to simulate time-dependent recordings, we first computed a
forward solution due to each epileptic source, assuming dipoles of unit-strength.
Each source produces a map of values at the simulated electrode
sites. Running forward simulations for each of several dipoles results
in a collection of several maps. To extend the single-instant values at
the electrodes into time-dependent signals, we scaled the values of each
map by pre-recorded clinical activation signals. Finally, we added ![]() noise to the projected data to better simulate physical EEG measurements.
noise to the projected data to better simulate physical EEG measurements.
The above method for solving the forward problem is needed not only to derive the simulated electrode recordings, but also as the iterative engine for solving the inverse source localization problem.
Next: Inverse Problem Up: Independent Component Analysis For Previous: Introduction Zhukov Leonid
Fri Oct 8 13:55:47 MDT 1999